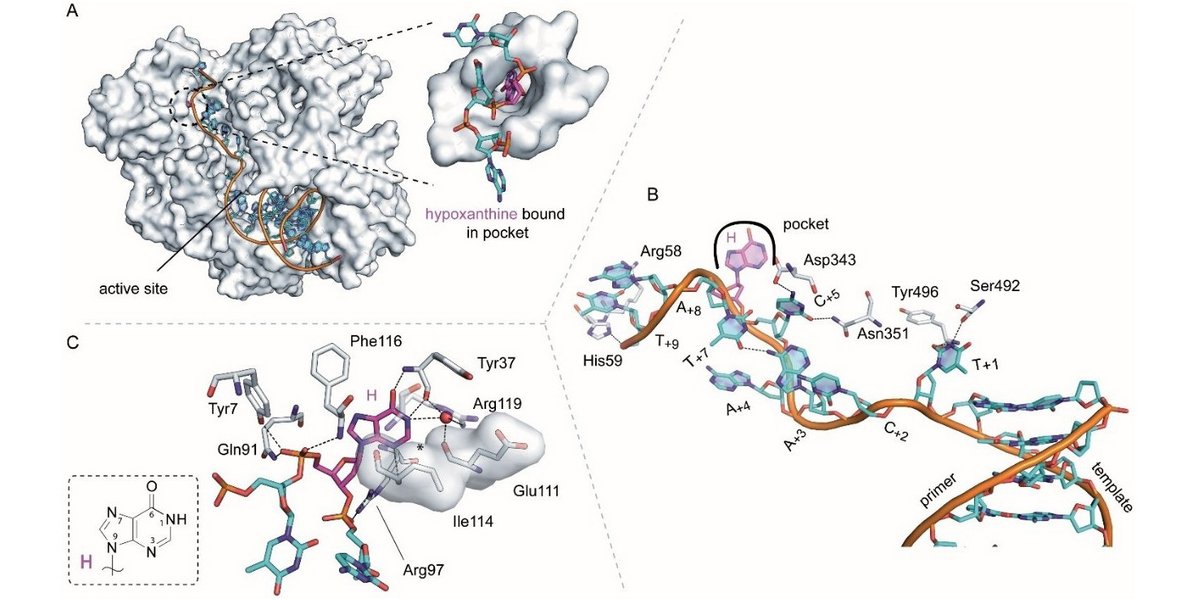
Structural Basis for The Recognition of Deaminated Nucleobases by An Archaeal DNA Polymerase
Heike M. Kropp, Samra Ludmann, Kay Diederichs, Karin Betz, and Andreas Marx (2021) ChemBioChem 22: pp 3060–3066
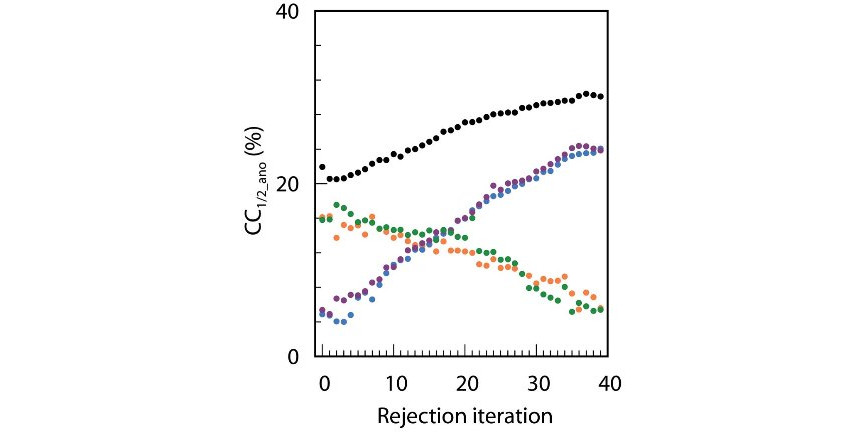
Making a difference in multi-data-set crystallography: simple and deterministic data-scaling/selection methods.
Assmann, G.M.,Wang, M., Diederichs, K. (2020) Acta Cryst D76, 636-552
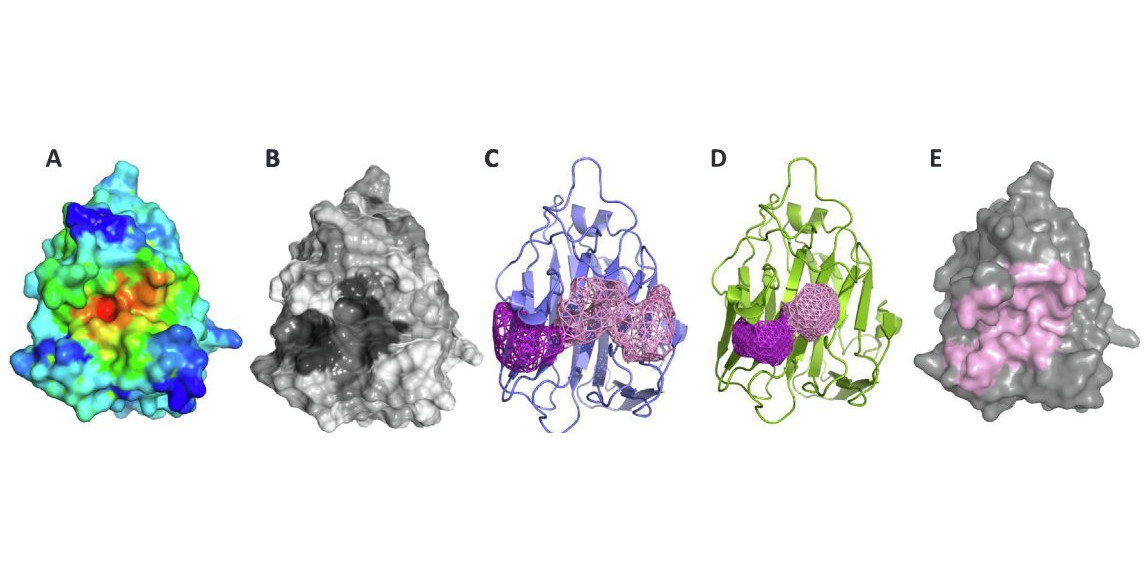
Structural annotation of the conserved carbohydrate esterase vb_24B_21 from Shiga toxin-encoding bacteriophage Φ24B
Barbara Franke, Marta Veses-Garcia, Kay Diederichs, Heather Allison, Daniel J. Rigden, and Olga Mayans (2020) Journal of Structural Biology 212: 107596
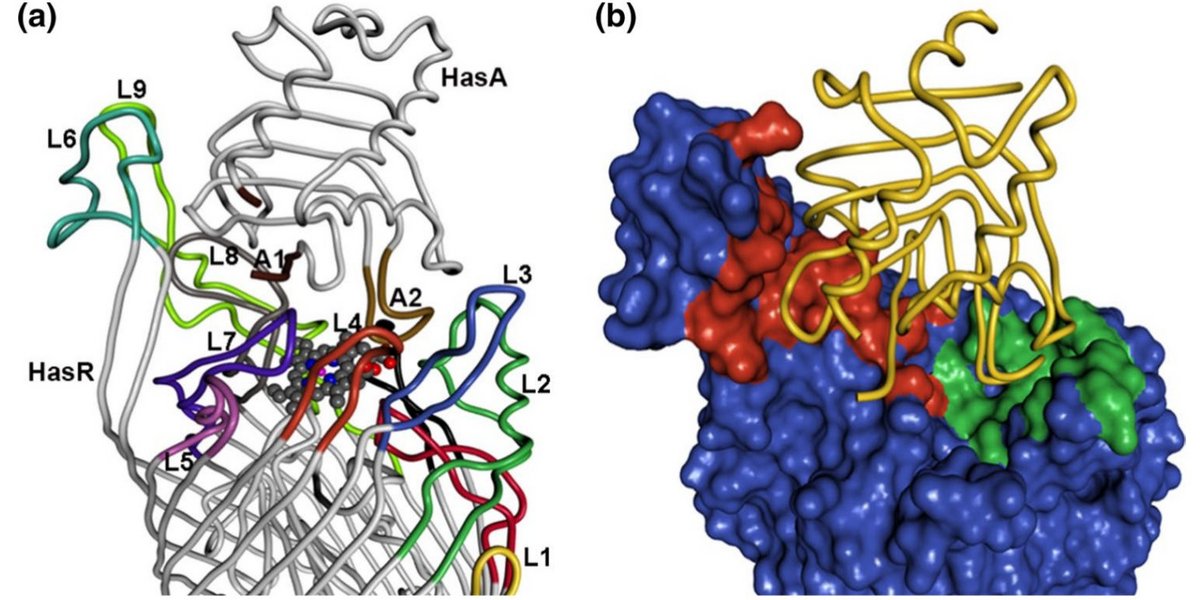
Binding of HasA by its transmembrane receptor HasR follows a conformational funnel mechanism.
Exner, T.E., Becker, S., Becker, S., Boniface-Guiraud, A., Delepelaire, P., Diederichs, K., Welte, W. (2020) Eur Biophys J. 49, pp 39-57
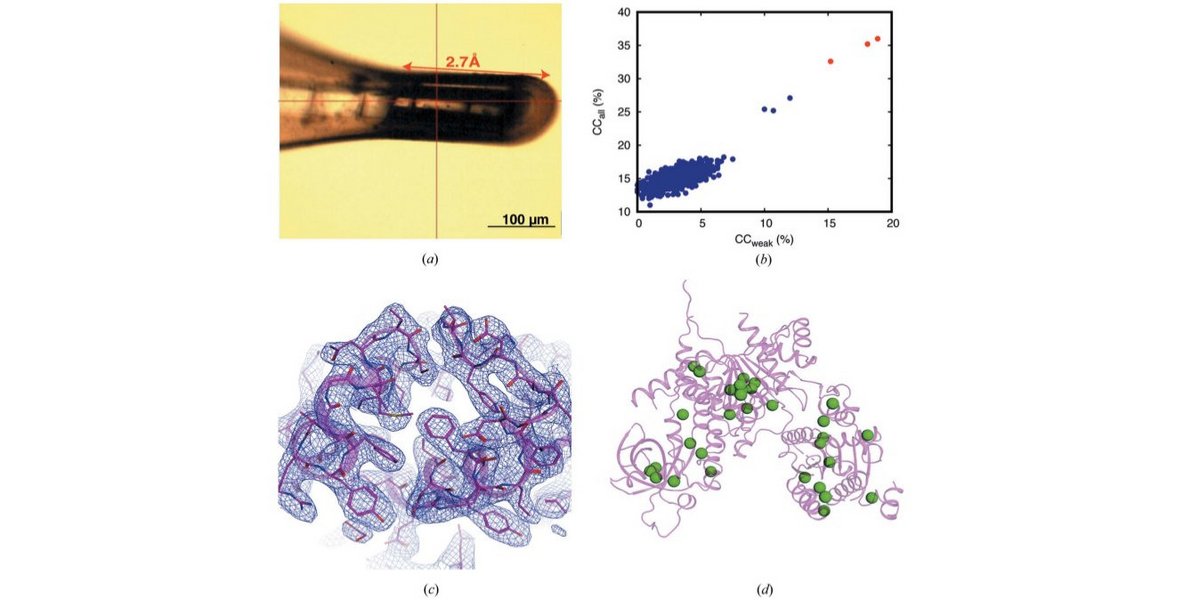
Long-wavelength native-SAD phasing: opportunities and challenges.
Basu, S., Olieric, V., Leonarski, F., Matsugaki, N., Kawano, Y., Takashi, T., Huang, C.Y., Yamada, Y., Vera, L., Olieric, N., Basquin, J., Wojdyla, J.A., Bunk, O., Diederichs, K., Yamamoto, M., Wang, M. (2019) JUCrJ 6, pp 373-386
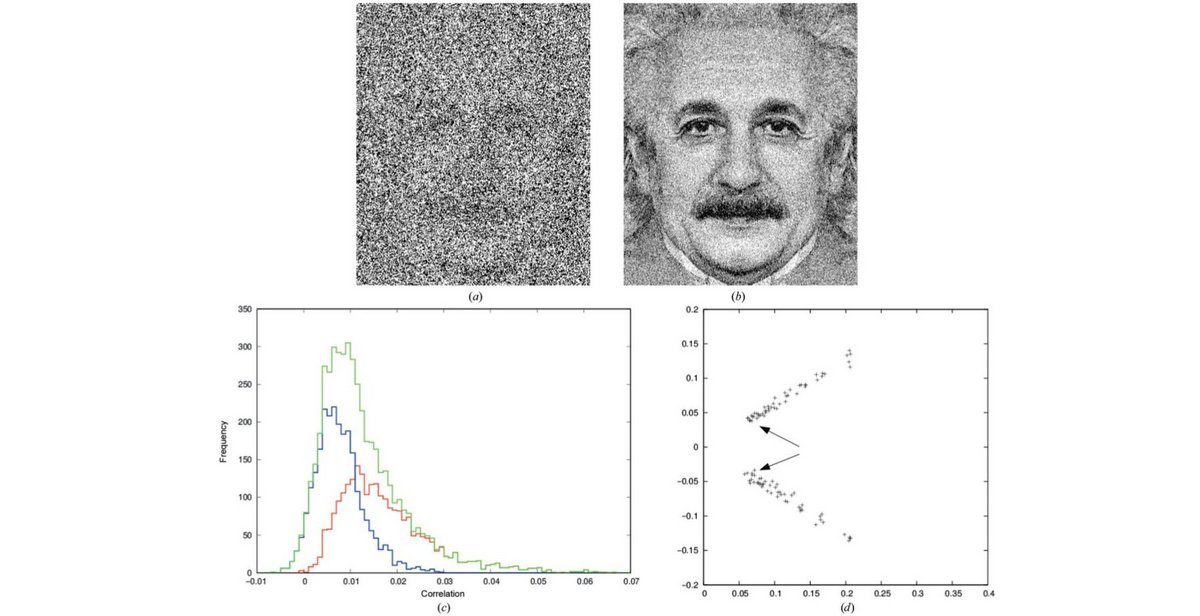
Dissecting random and systematic differences between noisy composite data sets.
K. Diederichs (2017) Acta Cryst D73, pp 286-293