Glycolytic pathways in Diatoms
Carolina Río Bártulos
Diatoms belong to the stramenopiles, a diverse group of organism which include photoautotrophic and heterotrophic species. Stramenopiles evolved by secondary endocytobiosis, the uptake of a photosynthetic eukaryote into a eukaryotic host cell. This evolutionary process increased the complexity of the resulting cells by combining two nuclear and three organellar genomes. Extensive gene transfers and genomic reorganization had strong implications on physiology and biochemistry of the resulting cells. Stramenopile genome projects reveal interesting distributions of metabolic pathways. One surprising finding is that all members of the stramenopiles investigated so far (including several diatom and oomycete species) possess nuclear encoded mitochondrial isoforms of enzymes of the second half of glycolysis, the oxidation of triosephosphate to pyruvate. This was unexpected as glycolysis in all other eukaryotes investigated so far, does not take place in the mitochondria. We also found a set of nucleus encoded glycolytic enzymes which seem to be target to the plastid in the investigated diatoms.
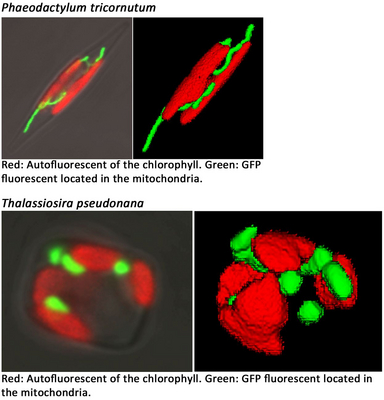
We verified this intracellular localization pattern of glycolytic enzymes in mitochondria, plastids and the cytosol via expression of GFP fusion proteins in the diatom Phaeodactylum tricornutum. Observation of P. tricornutum cells expressing GFP in the mitochondria furthermore revealed that mitochondria of this diatom show a high morphological variability. In order to study physiological functions we have established a protocol to isolate mitochondria from the diatom Thalassiosira pseudonana.