MAPP
Motif Activity on Pre-mRNA Processing
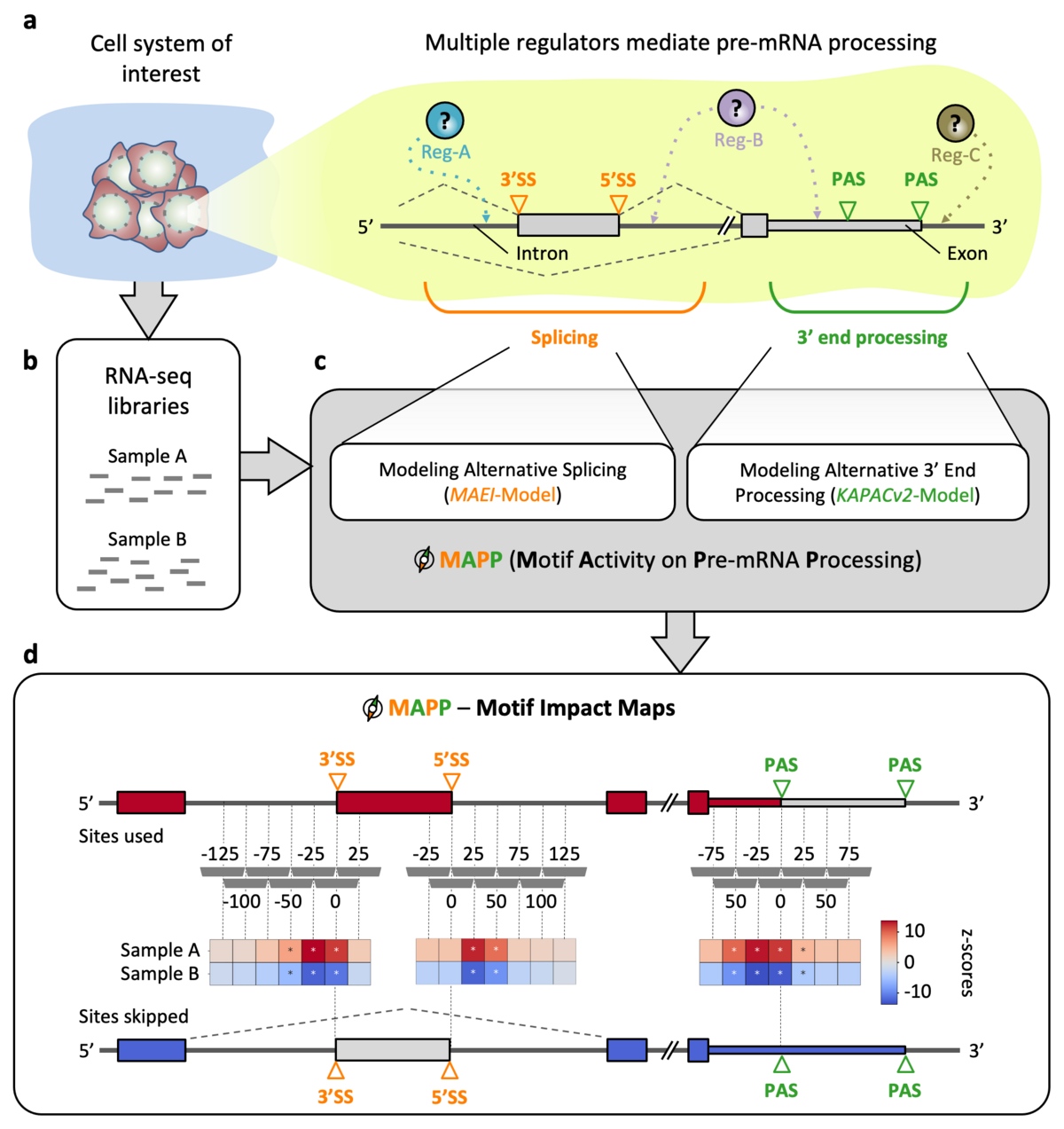
Maturation of eukaryotic precursor messenger RNAs (pre-mRNAs) via splicing, 3’ end cleavage and polyadenylation is modulated across cell types and conditions by a variety of RNA-binding proteins (RBPs). Both processes increase the variability of the transcriptome. Despite a significant number of proteins being associated with RNAs in human cells, their binding motifs as well as their functional impact is not fully understood. To characterize the impact of RBPs on splicing and/or polyadenylation (poly(A)) we have developed MAPP (Motif Activity on Pre-mRNA Processing), a fully automated computational tool. MAPP enables the inference of the RBP sequence motif-guided regulation of pre-mRNA processing from standard RNA sequencing (RNA-seq) data.
