News and announcements
Viaflo Assist plus helps to screen an enzyme library
Screening of a SP6 RNA polymerase library to identify a variant that accepts modified nucleotides.
AG Marx, Julia Gutbrod
Manual Assay with Vialfo Pipettes
Fluoresecence polarization based KD determination of mutant XNA polymerases in complex with unnatural substrates
AG Marx, Cédric Gutfreund
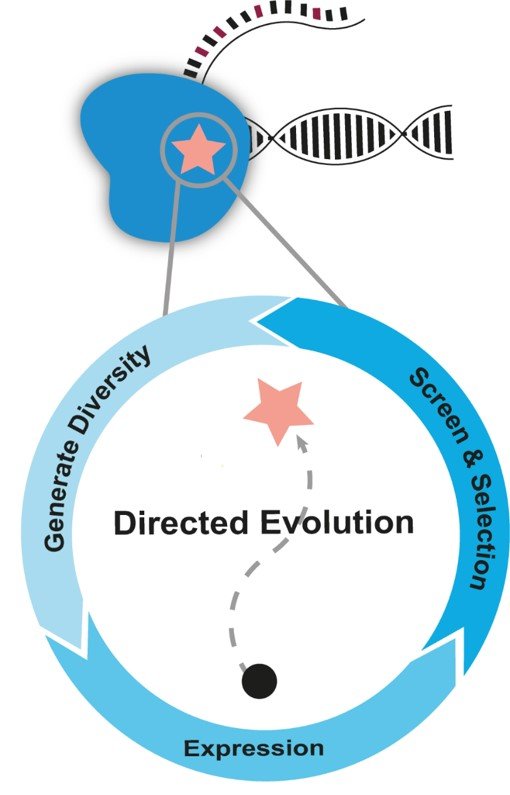
Viaflo Assist plus helps to screen an enzyme library
Screening of a T7 RNA polymerase library to identify a variant that accepts modified nucleotides.
AG Marx, Svenja Hehn
Screen to identify inhibitors of a novel RNA ligase
Screen to identify inhibitors of a novel RNA ligase (protein based screen, fluorescence)
AG Marx, Lisa Schlor
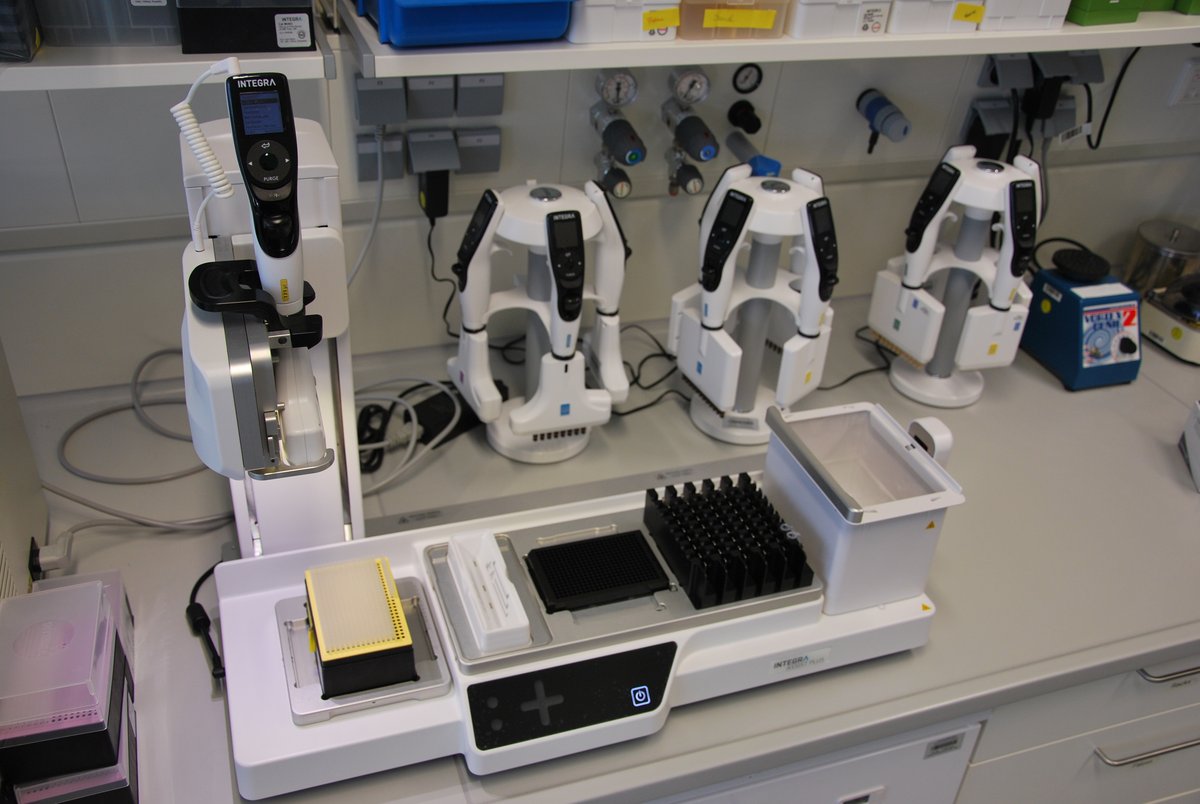

Viaflo Assit plus helps to identify candidates
Enginieering of a new Taq DNA polymerase variant with improved ability of reverse transcription
AG Marx, Luisa Huber
Screen to identify ubiquitin ligase mutant activators/inhibitors with our new natural substance library
Screen to identify ubiquitin ligase mutant activators (protein based screen, fluorescence polarization),
AG Martin Scheffner, Project Leader Franziska Müller
New compound library available
10.000 new natural compounds (in total now ~75.000 cpds)
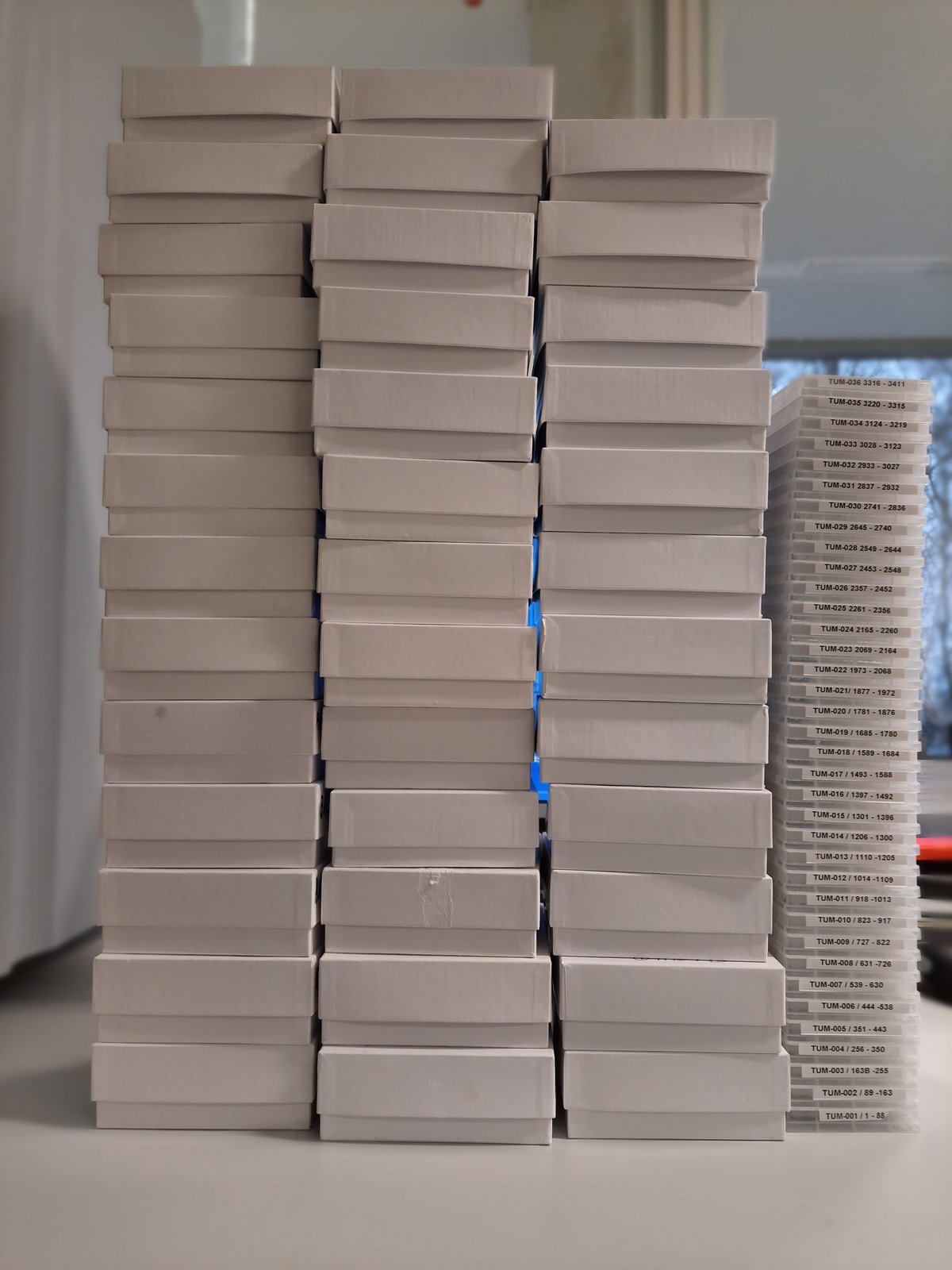

From much to small
Reformatting of a plasmidbank from eppendorf tubes to 96 well plate format
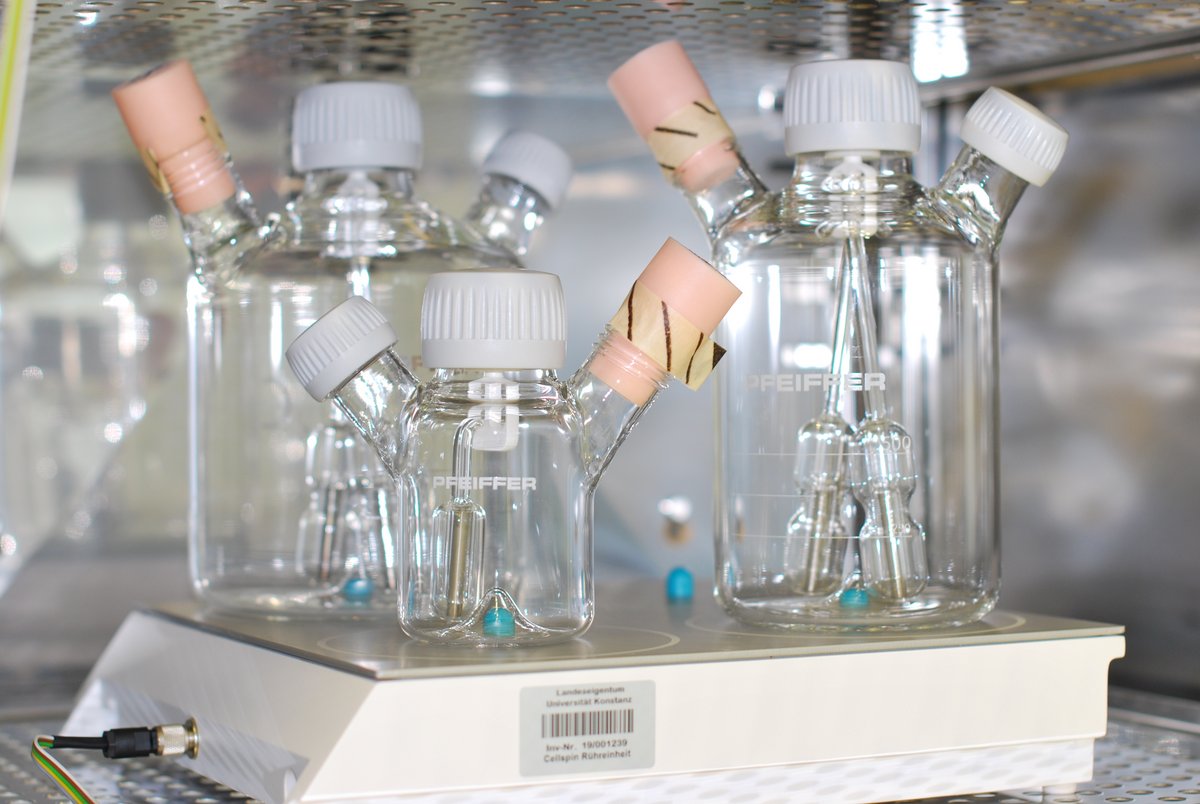

New service
Cell Culture Service for Hybridoma Antibody Production
New compound library available
700 new compounds with literature references
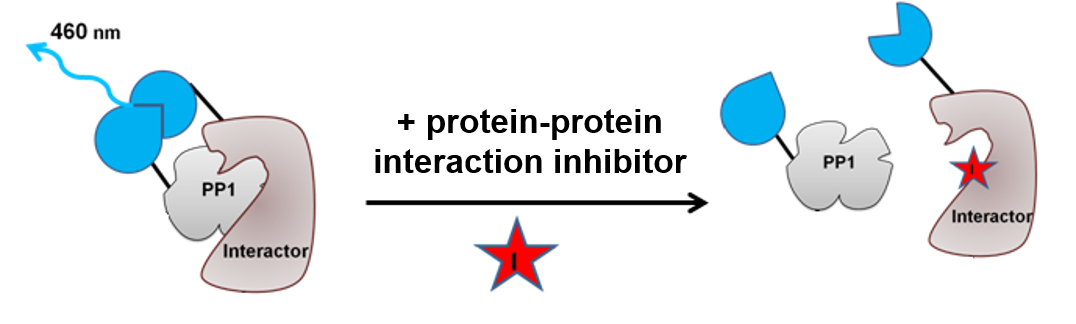
Screen for specific phosphatase holoenzyme inhibitors/activators
Split-luciferase based assay to identify protein-protein interaction inhibitors or stabilizers of phosphatase holoenzymes (protein based screen, luminescence),
Prof. Mathieu Bollen, Project Leader Zander Claes, KU Leuven
Screen to identify ubiquitin ligase mutant activators
Screen to identify ubiquitin ligase mutant activators (protein based screen, fluorescence polarization),
AG Martin Scheffner, Project Leader Franziska Müller
New drying method for compoundtransfertool
with help of our Werkstätten (thanks to them) a new holder for our fixed tips modul (compound transfer) could be developed. Drying the fixed tips is now more than 10% faster.
New software Scaffold Hunter for evaluation
We use Scaffold Hunter for Clustering and Visualisation of Compounds
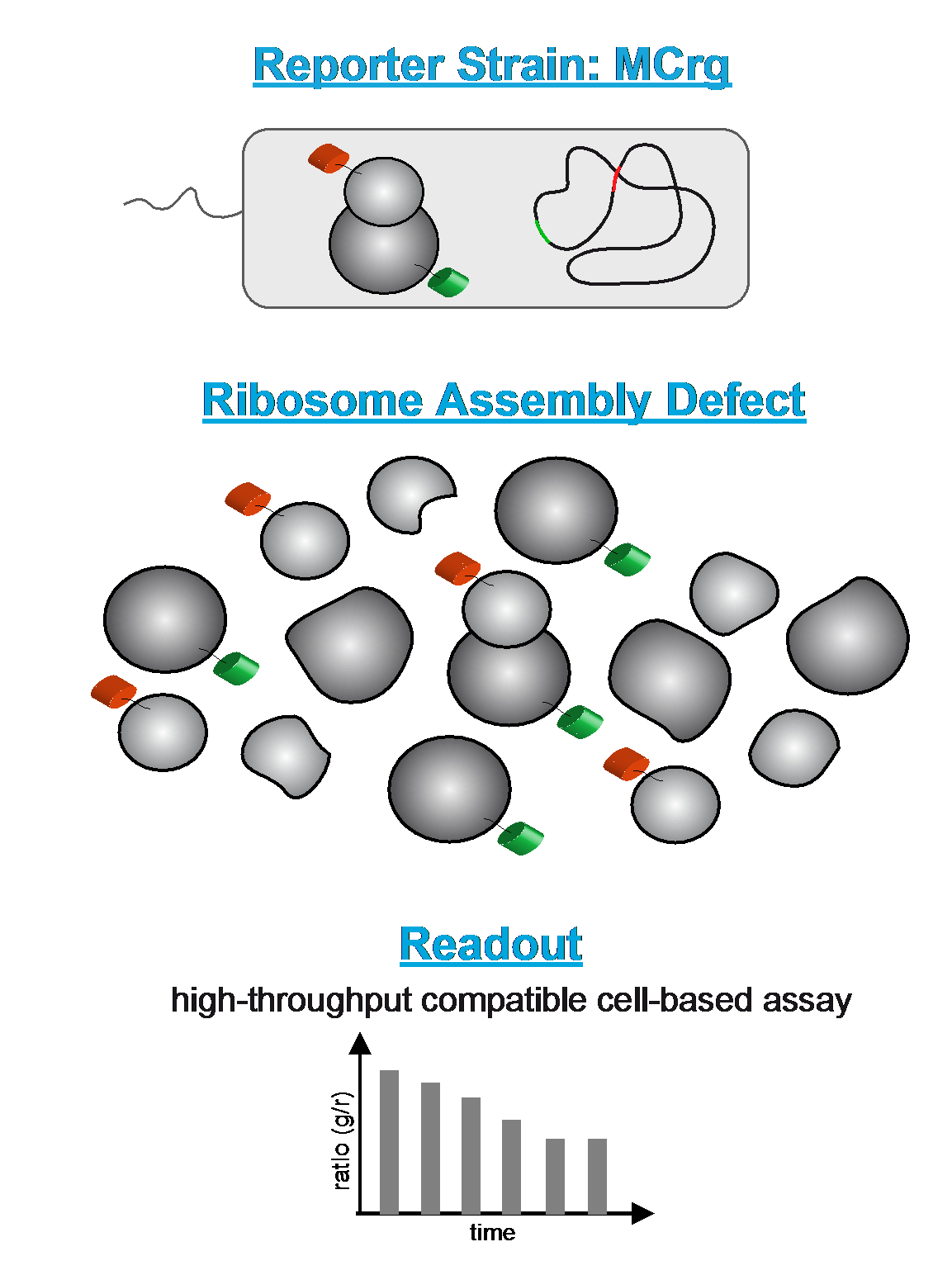
Re-Screen to identify factors involved in ribosomal subunit assembly
While the structure of mature ribosomes is analyzed in atomic detail, considerably less is known about their assembly process in living cells. This is mainly due to technical and conceptual hurdles. To facilitate the analysis of ribosome assembly in vivo, we designed fluorescently labeled Escherichia coli strains that allow the detection of ribosome assembly defects in a high-throughput compatible format. The assay is used to screen large small molecule libraries to identify factors involved in ribosomal subunit assembly. This should pave the way for a first identification of assembly inhibitors and thus potentially for the development of new classes of antimicrobial agents.
AG Deuerling, Project Leader Sabine Schmidt in Cooperation with Dr. Rainer Nikolay (Charité Berlin)
Screen for the identification of phosphatase effectors
Screen to identify human Ser/Thr phosphatase PPM1F inhibitors / activators by sensitive fluorogenic HTS. The particularity is a continuous readout of the enzyme reaction. (protein based screen, fluorescence)
AG Hauck, Project Leader Tanja Grimm
Assay optimization for the identification of phosphatase effectors
Screen to identify human Ser/Thr phosphatase PPM1F inhibitors / activators by sensitive fluorogenic HTS (protein based screen, fluorescence)
AG Hauck, Project Leader Tanja Grimm
Screen to identify ubiquitin ligase effectors
Screen to identify ubiquitin ligase effectors (protein based screen, fluorescence polarization),
AG Martin Scheffner, Project Leader Fabian Offensperger, Part 2
Counterscreen to part 1: Screen to identify ubiquitin ligase effectors
Counterscreen to part 1: Screen to identify ubiquitin ligase effectors (protein based screen, fluorescence polarization),
AG Martin Scheffner, Project Leader Fabian Offensperger
Screening for RNA binding molecules
Screening for RNA binding molecules (e.coli based screen, luminescence),
AG Hartig, Project Leader Malte Sinn
New compound library available
new compound library available (July 2016): in total now more than 60.000cpds
Screen to identify ubiquitin ligase effectors
Screen to identify ubiquitin ligase effectors (protein based screen, fluorescence polarization),
AG Martin Scheffner, Project Leader Fabian Offensperger, Part 1

Restart after Optimization: High Content Screen to identify Separase Inhibitors
Restart after Optimization: High Content Screen to identify Separase Inhibitors (human cell based screen),
AG Thomas U. Mayer (Uni KN) / AG Stemmann (Uni Bayreuth), Lars Henschke
Screening Database completed
Screening Database completed: all compounds with structure in
Two new compound libraries available
Two new compound libraries available: in total now ~ 55.000cpds
